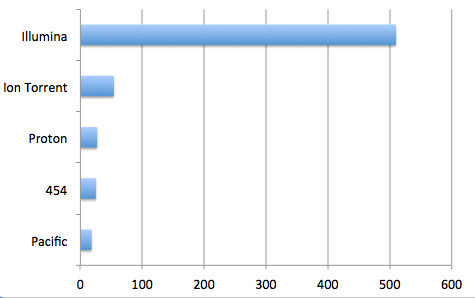
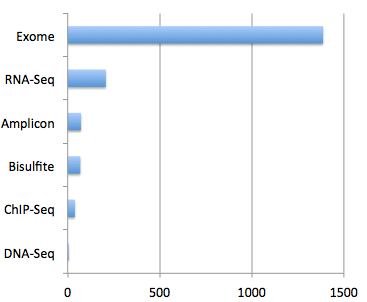
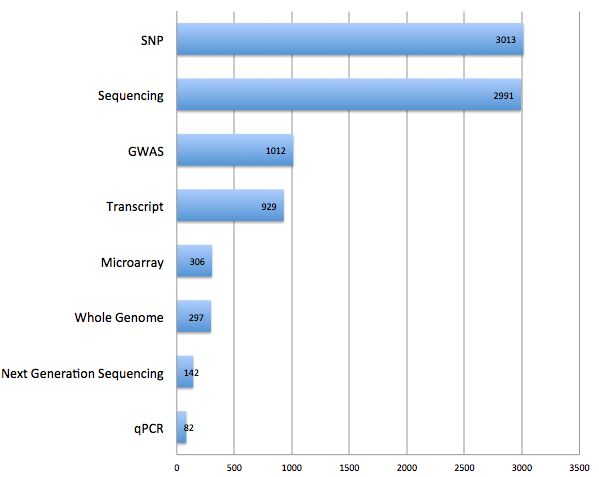
The ability to list and purchase bioinformatic analysis services along with next-gen sequencing comes to Genohub. This additional feature set will further simplify the NGS project experience by offering capabilities for the bioinformatics project step, all in one roof. By launching this service, researchers on Genohub are now able to shop from full-service sequencing providers who offer the three main services required in every NGS project with options for Illumina, Ion, SOLiD, PacBio, and Roche454 platforms:
- Library prep
- Next generation sequencing
- Bioinformatic analysis
The addition of this service alleviates the burden many researchers face of shopping for three separate sequencing related services from multiple providers and efficiently managing the entire process.
The benefits of this feature for researchers includes:
- Researchers save time and money purchasing sequencing and bioinformatic services simultaneously
- By enabling next gen sequencing providers to list bioinformatic services, it becomes easier and faster for researchers to compare and select a provider for their project
- Risk is mitigated, as researchers are able to purchase bioinformatic services along with NGS services by providers that have gone through the Genohub vetting process
Providers also benefit:
- Providers are now able to better highlight their unique services by listing specific bioinformatic services. These include basic primary analysis that may be included with every sequencing order, as well as more complex secondary and custom bioinformatics services that can be ordered as add-ons to NGS projects.
- The bioinformatics platform enables providers to more succinctly communicate bioinformatic services to researchers, cutting down on back and forth communication prior to the researcher opting to move forward with the services listed by the provider
Providers may now list bioinformatic analysis services on the familiar “Manage Services Offerings” page:
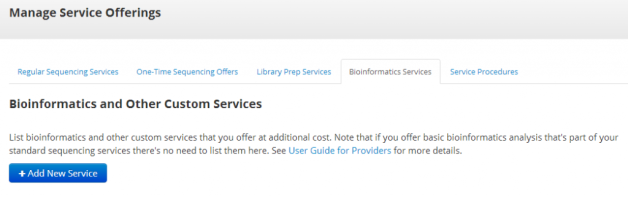
New Manage Services Page
Researchers will be able to view these services once they select to view details about a sequencing package found when shopping by project or technology.
At Genohub we constantly strive to improve the service we offer to the NGS community through the development of new features and functionality. When we launched Genohub in August, 2013 our goal was to launch a service that was useful to researchers and sequencing providers, and continue to enhance and improve our service based directly on feedback from our clients.
The bioinformatic analysis listing feature come as a direct result of our development methodology. Researchers asked for a one-stop-shop for all of their sequencing needs, and sequencing providers asked for more ways to highlight and differentiate their services on Genohub. You asked, we listened!
Please visit Genohub often as we are constantly adding new useful features to improve the next-gen sequencing shopping experience including upcoming features to further enable providers to differentiate themselves and mitigate risk in decision making for researchers. If you have any feedback on additional features you would like to see, please send us feedback by shooting us an email at info@genohub.com.